AIPMT 2012 Prelims
Paper was held on
Sun, Apr 1, 2012 10:00 AM
Biology
1
Which one of the following pairs of hormones are the examples of those that can easily pass through the cell membrane of the target cell and bind to a receptor inside it (mostly in the nucleus)?
2
During gamete formation, the enzyme recombinase participates during
3
A certain road accident patient with unknown blood group needs immediate blood transfusion. His one doctor friend at once offers his blood. What was the blood group of the donar?
4
People who have migrated from the planes to an area adjoining Rohtang Pass about six months back
5
Which one of the following is the correct statement for respiration in humans?
6
Select the correct statement regarding the specific disorder of muscular or skeletal system.
7
Which part of the human ear plays no role in hearing as such but is otherwise very much required?
8
The human hind brain comprises three parts, one of which is
9
The maximum amount of electrolytes and water (70 $$-$$ 80 percent) from the glomerular filtrate is reabsorbed in which part of the nephron?
10
What is correct to say about the hormone action in humans?
11
A person entering an empty room suddenly finds a snake right in front on opening the door. Which one of the following is likely to happen in his neuro-hormonal control system?
12
In a normal pregnant woman, the amount of total gonadotropin activity was assessed. The results
expected was :
13
An organic substance that can withstand environmental extremes and cannot be degraded by any enzyme is
14
Signals for parturition originate from :
15
Which one of the following statements is false in respect of viability of mammalian sperm?
16
Even in absence of pollinating agents seed-setting is assured in
17
Both, autogamy and geitonogamy are prevented in
18
Which one of the following is correctly matched?
19
The leydig's cells as found in the human body are the secretory source of
20
What is the figure given below showing in particular ?


21
The test-tube baby programme employs which one of the following techniques ?
22
A normal - visioned man whose father was colour bilind, marries a woman whose father was also
colour - blind. They have their first child as a daughter. What are the chances that this child would be
colour blind
23
The gynoecium consists of many free pistils in flowers of
24
The most abundant prokaryotes helpful to humans in making curd ftom milk and in production of antibiotics are the ones categorised as
25
Cycas and Adiantum resemble each other in having
26
Which one single organism or the pair of organisms is correctly assigned to its or their named taxonomic group?
27
Which one of the following is a correct statement?
28
The cyanobacteria are also referred to as
29
Maximum nutritional diversity is found in the group
30
Which one of the following is common to multicellular fungi, filamentous algae and protonema of mosses?
31
Nuclear membrane is absent in
32
Which one of the following microbes forms symbiotic association with plants and helps them in their nutrition?
33
In which one of the following, the genus name, its two characters and its phylum are not correctly matched, whereas the remaining three are correct?
34
Pheretima and its close relatives derive nourishment from
35
Cymose inflorescence is present in
36
F2 generation in a Mendelian cross showed that both genotypic and phenotypic ratios are same as 1 : 2
: 1. It represents a case of
37
How many plants in the list given below have composite fruits that develop from an inflorescence?
Walnut, poppy, radish, fig, pineapple, apple, tomato, mulberry.
Walnut, poppy, radish, fig, pineapple, apple, tomato, mulberry.
38
Phyllode is present in
39
Placentation in tomato and lemon is
40
Vexillary aestivation is characteristic of the family
41
The coconut water and the edible part of coconut are equivalent to
42
The given diagrammatic representation shows one of the caregories of small molecular weight organic compounds in the living tissues. Identify the category shown and the one blank component ''X'' in it.


43
Which one out of A - D given below correctly represents the structural formula of the basic amino acid?


44
Which one of the following is wrong statement?
45
Which one is the most abundant protein in the animal world?
46
The correct sequence of cell organelles during photores piration is
47
A process that makes important difference between C3 and C4 plants is
48
Which one of the following is not a functional unit of an ecosystem?
49
Which one of the following is not a gaseous biogeochemical cycle in ecosystem ?
50
Giiven below is an imaginary pyramid of numbers. What could be one of the possibilities about certain
organisms at some of the different levels ?


51
The upright pyramid of number is absent in
52
Identify the possible link "A" in the following food chain :
Plant → insect → frog →"A"→ Eagle
Plant → insect → frog →"A"→ Eagle
53
Which one of the following areas in India, is a hotspot of biodiversity?
54
The common bottle cork is a product of
55
Water containing cavities in vascular bundles are found in
56
Gymnosperms are also called soft wood spermatophytes because they lack
57
Closed vascular bundles lack
58
Companion cells are closely associated with
59
For transformation, micro - particles coated with DNA to be bombarded with gene are made up of
60
Which one of the following does not differ in E.coli and Chlamydomonas ?
61
Select the the correct statement from the following regarding cell membrane
62
Compared to those of humans, the erythrocytes in frog are :
63
Select the correct statement from the ones given below with respect to Periplaneta americana
64
What is true about ribosomes ?
65
Widal Test is carried out to test :
66
In which one of the following options the two
examples are correctly matched with their
particular type of immunity?
67
Given below is the representation of a certain event at a particular stage of a type of cell division. Which is this stage?


68
The highest number of species in the world
is represented by
69
Yeast is used in the production of
70
Cirrhosis of liver is caused by the chronic intake of
71
Which one of the following is not a part of a transcription unit in DNA ?
72
Removal of introns and joining of exons in a defined order during transcription is called
73
If one strand of DNA has the nitrogenous base sequence as ATCTG, what would be the
complementary RNA stand sequence
74
Removal of RNA polymerase -III from nucleoplasm will affect the synthesis of
75
Ribosomal RNA is actively synthesized in
76
Which one of the following options gives one correct example each of convergent evolution and
divergent evolution ?
77
What was the most significant trend in the evolution of modern man (Homo sapiens) from his
ancestors ?
78
Evolution of different species in a given area starting from a point and spreading to other
geographical areas is known as
79
The extinct human who lived 1,00,000 to 40,000 years ago, in Europe, Asia and parts of Africa, with
short stature, heavy eye brows, retreating fore heads, large jaws with heavy teeth, stocky bodies, a
lumbering gait and stooped posture was :
80
Which one of the following is a case of wrong matching ?
81
A nitrogen - fixing microbe associated with Azolla in rice fields is
82
Motile zygote of Plasmodium occurs in
83
Which statement is wrong for viruses?
84
A patient brought to a hospital with myocardial infarction is normally immediately given
85
Common cold differs from pneumonia in, that
86
Which part would be most suitable for raising virus free plants for microporpagation ?
87
Which one of the following is an example of carrying out biological control of pests/diseases using
microbes ?
88
Which one of the following in not a property of cancerous cells whereas the remaining three are ?
89
Monascus purpureus is a yeast used commercially in the production of :
90
The figure below is the diagrammatic respesentation o the E.Coli vector pBR 322. Which one of the
given options correctly identifies its certain component(s) ?


91
PCR and Restriction Fragment Length Polymorphism are the methods for
92
Which one is a true statement regarding DNA polymerase used in PCR ?
93
A single strand of nucleic acid tagged with a radioactive molecule is called
94
Consumption of which one of the following foods can prevent the kind of blindness associated with
vitamin 'A' deficiency
Chemistry
1
Maximum number of electrons in a subshell with $$l$$ = 3 and n = 4 is
2
CH3CHO and C6H5CH2CHO can be distinguished chemically by
3
Deficiency of vitamin B1 causes the disease
4
Which one of the following sets of monosaccharides forms sucrose?
5
pA and pB are the vapour pressure of pure liquid components, A and B, respectively of an ideal binary solution. If xA represents the mole fraction of component A, the total pressure of the solution will be
6
In a zero-order reaction, for every 10oC rise of temperature, the rate is doubled. If the temperature is increased from 10oC to 100oC, the rate of the reaction will become
7
Limiting molar conductivity of NH4OH
$$\left[ {} \right.$$i.e. $$\Lambda _{m\left( {N{H_4}OH} \right)}^0$$$$\left. {} \right]$$ is equal to
$$\left[ {} \right.$$i.e. $$\Lambda _{m\left( {N{H_4}OH} \right)}^0$$$$\left. {} \right]$$ is equal to
8
In a reaction, A + B $$ \to $$ product, rate is doubled when the concentration of B is doubled, and rate increases by a factor of 8 when the concentration of both the reactants (A and B) are doubled, rate law for the reaction can be written as
9
In which of the following compounds, nitrogen exhibits highest oxidation state?
10
A mixture of potassium chlorate, oxalic acid and sulphuric acid is heated. During the reaction which element undergoes maximum change in the oxidation number?
11
Which of the following statements is not valid for oxoacids of phosphorus?
12
Sulphur trioxide can be obtained by which of the following reaction?
13
Which of the statements is not true?
14
Identify the alloy containing a non-metal as a constituent in it.
15
Which one of the following is an outer orbital complex and exhibits paramagnetic behaviour?
16
Which nomenclature is not according to IUPAC system?
17
Identify the wrong statement in the following.
18
The correct set of four quantum numbers for the valence electron of rubidium atom (Z = 37) is
19
Which one of the following pairs is isostructural (i.e., having the same shape and hybridization) ?
20
The pair of species with the same bond order is
21
Bond order of 1.5 is shown by
22
Which of the following species contains three bond pairs and one lone pair arround the central atom ?
23
In which of the following reactions, standard reaction entropy change ($$\Delta $$So) is positive and standard Gibb's energy change ($$\Delta $$Go) decreases sharply with increasing temperature ?
24
Standard enthalpy of vaporisation $$\Delta $$vapHo for water at 100oC is 40.66 kJ mol$$-$$1. The internal energy of vaporisation of water at 100oC (in kJ mol$$-$$1) is
25
The enthalpy of fusion of water is 1.435 kcal/mol. The molar entropy change for the melting of ice at 0oC is
26
pH of a saturated solution of Ba(OH)2 is 12. The value of solubility product (Ksp) of Ba(OH)2 is
27
Buffer solutions have constant acidity and alkalinity because
28
Among the following compounds the one that is most reactive towards electrophilic nitration is
29
Equimolar solutions of the following substances were prepared separately. Which one of these will record the highest pH value ?
30
Which of the following acids does not exhibit optical isomerism?
31
In the following reaction

The major product is

The major product is
32
In the following sequence of reactions

the end product (C) is

the end product (C) is
33
When Cl2 gas reacts with hot and concentrated sodium hydroxide solution, the oxidation number of chlorine changes from
34
Acetone is treated with excess of ethanol in the presence of hydrochloric acid. The product obtained is
35
The correct order of decreasing acid strength of trichloroacetic acid (A), trifluoroacetic acid (B), acetic acid (C) and formic acid (D) is
36
Predict the products in the given reaction.


Physics
1
A ray of light is incident at an angle of incidence i, on one face of a prism of angle A (assumed to be small ) and emerges normally from the opposite face. If the refractive index of the prism is $$\mu $$. the angle of incidence i, is nearly equal to
2
A ring is made of a wire having a resistance R0 = 12 $$\Omega $$. Find the points A and B, as shown in the figure, at which a current carrying conductor should be connected so that the resistance R of the sub circuit between these point is equal to $${8 \over 3}\Omega $$.
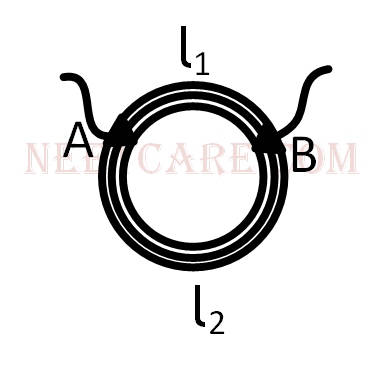

3
A compass needle which is allowed to move in a horizontal plane is taken to a geomagnetic pole. It
4
Four point charges $$-$$Q, $$-$$q, 2q and 2Q are placed, one at each corner of the square. The relation between Q and q for which the potential at the centre of the square is zero is
5
If voltage across a bulb rated 220 volt-100 watt drops by 2.5% of its rated value, the percentage of the rated value by which the power would decrease is
6
In an electrical circuit R, L, C and a.c. voltage source are all connected in series. When L is removed from the circuit, the phase difference between the voltage and the current in the circuit is If instead, C is removed from the circuit, the phase difference is again. The power factor of the circuit is
7
The current $$(I)$$ in the inductance is varying with time according to the plot shown in figure.

Which one of the following is the correct variation of voltage with time in the coil ?

Which one of the following is the correct variation of voltage with time in the coil ?
8
A coil of resistance 400 $$\Omega $$ is placed in a magnetic field. If the magnetic flux $$\phi $$ (Wb) linked with the coil varies with time t (sec) as $$\phi = 50{t^2} + 4$$.
The current in the coil at t = 2 sec is
The current in the coil at t = 2 sec is
9
The electric field associated with an em wave in vacuum is given by
E = $$\widehat i$$ 40 cos (kz $$-$$ 6 $$ \times $$ 108 t) where E, z and t are in volt/m, meter and seconds respectively. The value of wave vector k is
E = $$\widehat i$$ 40 cos (kz $$-$$ 6 $$ \times $$ 108 t) where E, z and t are in volt/m, meter and seconds respectively. The value of wave vector k is
10
A 200 W sodium street lamp emits yellow light of wavelength 0.6 $$\mu $$m. Assuming it to be 25% efficient in converting electrical energy to light, the number of photons of yellow light it emits per second is
11
An $$\alpha $$-particle moves in a circular path of radius 0.83 cm in the presence of a magnetic field of 0.25 Wb/m2. The de Broglie wavelength associated with the particle will be
12
Monochromatic radiation emitted when electron on hydrogen atom jumps from first excited to the ground state irradiates a photosensitive material. The stopping potential is measured to be 3.57 V. The threshold frequency of the material is
13
The magnifying power of a telescope is 9. When it is adjusted for parallel rays the distance between the objective and eyepiece is 20 cm. The focal length of lenses are
14
Two similar coils of radius R are lying concentrically with their planes at right angles to each other. The currents flowing in them are $$I$$ and 2$$I$$, respectively. The resultant magnetic field induction at the centre will be
15
When a biconvex lens of glass having refractive index 1.47 is dipped in a liquid, it acts as a plane sheet of glass. This implies that the liquid must have refractive index
16
A concave mirror of focal length $$f$$1 is placed at a distance of d from a convex lens of focal length $$f$$2. A beam of light coming from infinity and falling on this convex lens $$-$$ concave mirror combination returns to infinity. The distance d must equal
17
Electron in hydrogen atom first jumps from third excited state to second excited state and then from second excited to the first excited state. The ratio of the wavelengths $$\lambda $$1 : $$\lambda $$2 emitted in the two cases is
18
An electron of a stationary hydrogen atom passes from the fifth energy level to the ground level. The velocity that the atom acquired as a result of photon emission will be
19
The figure shows a logic circuit with two inputs A and B and the output C. The voltage wave forms across A, B and C are as given. The logic circuit gate is


20
Two ideal diodes are connected to a battery as shown in the circuit. The current supplied by the battery is


21
C and Si both have same lattice structure; having 4 bonding electrons in each. However, C is insulator where as Si is intrinsic semiconductor. This is because
22
A spherical planet has a mass MP and diameter DP. A particle of mass m falling freely near the surface of this planet will experience an acceleration due to gravity, equal to
23
The motion of a particle along a straight line is described by equation x = 8 + 12t $$-$$ t3 where x is in metre and t in second. The retardation of the particle when its velocity becomes zero is
24
Two spheres A and B of masses m1 and m2 respectively collide. A is at rest initially and B is moving with velocity v along x-axis. After collision B has a velocity $${v \over 2}$$ in a direction perpendicular to the original direction. The mass A moves after collision in the direction :
25
The horizontal range and the maximum height of a projectile are equal. The angle of projection of the projectile is
26
The potential energy of a particle in a force field is $$U = {A \over {{r^2}}} - {B \over r}$$ where A and B are positive constants and r is the distance of particle from the centre of the field. For stable equilibrium, the distance of the particle is
27
A particle has initial velocity $$\left( {2\overrightarrow i + 3\overrightarrow j } \right)$$ and acceleration $$\left( {0.3\overrightarrow i + 0.2\overrightarrow j } \right)$$. The magnitude of velocity after 10 seconds will be
28
A car of mass 1000 kg negotiates a banked curve of radius 90 m on a frictionless road. If the banking angle is 45o, the speed of the car is
29
A solid cylinder of mass 3 kg is rolling on a horizontal surface with velocity 4 m s$$-$$1. It collides with a horizontal spring of force constant 200 N m$$-$$1. The maximum compression produced in the spring will be
30
When a mass is rotating in a plane about a fixed point, its angular momentum is directed along
31
ABC is an equilateral triangle with O as its centre. $${\overrightarrow F _1},{\overrightarrow F _2}$$ and $${\overrightarrow F _3}$$ represent three forces acting along the sides AB, BC and AC respectively. If the total torque about O is zero then magnitude of $${\overrightarrow F _3}$$ is


32
Two persons of masses 55 kg and 65 kg respectively, are at the opposite ends of a boat. The length of the boat is 3.0 m and weights 100 kg. The 55 kg man walks up to the 65 kg man and sits with him. If the boat is in still water the centre of mass of the system shifts by :
33
Liquid oxygen at 50 K is heated to 300 K at constant pressure of 1 atm. The rate of heating is constant. Which one of the following graphs represents the variation of temperature with time ?
34
The height at which the weight of a body becomes $${\left( {{1 \over {16}}} \right)^{th}}$$, its weight on the surface of earth (radius R), is
35
If the radius of a star is R and it acts as a black body, what would be the temperature of the star, in which the rate of energy production is Q?
36
A geostationary satellite is orbiting the earth at a height of 5R above the surface of the earth, R being the radius of the earth. The time period of another satellite in hours at a height of 2R from the surface of the earth is
37
A thermodynamic system is taken through the cycle ABCD as shown in figure. Heat rejected by the gas during the cycle is


38
One mole of an ideal gas goes from an initial state A to final state B via two processes : It first undergoes isothermal expansion from volume V to 3V and then its volume is reduced from 3V to V at constant pressure. The correct P-V diagram representing the two process is
39
Two sources of sound placed close to each other, are emitting progressive waves given by
y1 = 4sin600$$\pi $$t and y2 = 5sin608$$\pi $$t
An observer located near these two sources of sound will hear
y1 = 4sin600$$\pi $$t and y2 = 5sin608$$\pi $$t
An observer located near these two sources of sound will hear
40
When a string is divided into three segments of length $$l$$1, $$l$$2 and $$l$$3 the fundamental frequencies of these three segments are $${\upsilon _1},{\upsilon _2}$$ and $${\upsilon _3}$$ respectively. The original fundamental frequency ($$v$$) of the string is
41
A milli voltmeter of 25 milli volt range is to be converted into an ammeter of 25 ampare range. The value (in ohm) of neccessary shunt will be
42
An alternating electric field, of frequency $$v$$, is applied across the does (radius = R) of a cyclotron that is being used to accelerate protons (mass = m). The operating magnetic field (B) used in the cyclotron and the kinetic energy (K) of the proton beam, produced by it, are given by
43
What is the flux through a cube of side $$a$$ if a point charge of q is at one of its corner?
44
An electric dipole of moment p is placed in an electric field of intensity E. The dipole acquires a position such that the axis of the dipole makes an angle $$\theta $$ with the direction of the field. Assuming that the potential energy of the dipole to be zero when $$\theta $$ = 90o, the torque and the potential energy of the dipole will respectively be
45
In the circuit shown the cells A and B have negligible resistances. For VA = 12 V, R1 = 500 $$\Omega $$ and R = 100 $$\Omega $$ the galvanometer (G) shows no deflection. The value of Vs is
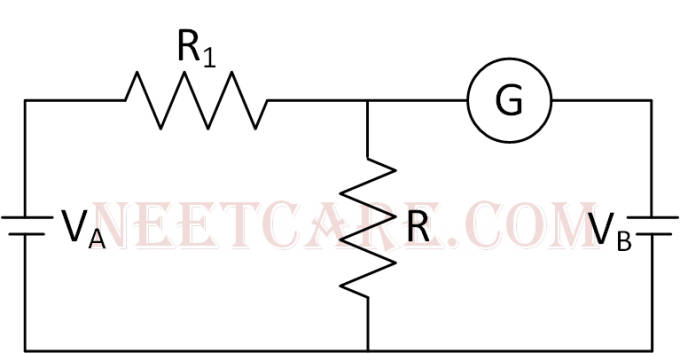

46
The damping force on an oscillator is directly proportional to the velocity. The units of the constant of proportionality are